Multimedia
Filter :
Publication date
Article type
Subject

Drop Impact on a Solid Surface: Supplemental Video 9
A supplemental video from the 2016 review by C. Josserand and S.T. Thoroddsen "Drop Impact on a Solid Surface" from the Annual Review of Fluid Mechanics.
Shown: Ejected droplets and azimuthal undulations on the lamellar surface. Frame rate is 125 kfps. From Thoroddsen et al. (2012).
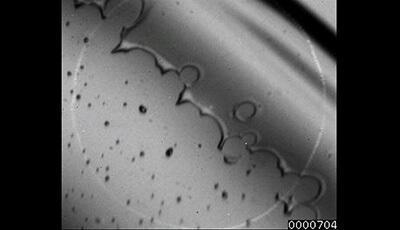

Drop Impact on a Solid Surface: Supplemental Video 10
A supplemental video from the 2016 review by C. Josserand and S.T. Thoroddsen "Drop Impact on a Solid Surface" from the Annual Review of Fluid Mechanics.
Shown: Air entrapment under a levitated lamella resulting from local contacts ahead of the moving contact line. Frame rate is 62 kfps. From Thoroddsen et al. (2010).

Drop Impact on a Solid Surface: Supplemental Video 11
A supplemental video from the 2016 review by C. Josserand and S.T. Thoroddsen "Drop Impact on a Solid Surface" from the Annual Review of Fluid Mechanics.
Shown: The impact of a compound drop outer drop is a water/glycerin mixture of viscosity 2 cP. It contains 20 inner perfluorohexane droplets which are more dense and migrate toward the bottom free surface of the large drop. For impact velocity V ≃ 4 m/s drop D ≃ 4 mm. Frame rate is 12 kfps. Courtesy of Jiaming Zhang and Erqiang Li.
Bacterial Hydrodynamics: Supplemental Video 1
A supplemental video from the 2016 review by Eric Lauga "Bacterial Hydrodynamics" from the Annual Review of Fluid Dynamics.
Shown: Swimming Escherichia coli bacteria with fluorescently labeled flagellar filaments showing fluorescent bundles of flagellar filaments rotating and propelling individual cells forward. The details of the experimental procedure are given in Turner L Ryu WS Berg HC. 2000. Real-time imaging of fluorescent flagellar filaments. J. Bacteriol. 182:2793–801. The video is reproduced with permission from Howard Berg’s website at the Rowland Insitute Harvard University (http://www.rowland.harvard.edu/labs/bacteria/movies/ecoli.php).
Bacterial Hydrodynamics: Supplemental Video 2
A supplemental video from the 2016 review by Eric Lauga "Bacterial Hydrodynamics" from the Annual Review of Fluid Dynamics.
Shown: Swimming Escherichia coli bacteria with fluorescently-labeled flagellar filaments showing individual run-and-tumble events with flagellar filaments unbundling cells turning and flagellar filaments rejoining the bundle. The details of the experimental procedure are given in Turner L Ryu WS Berg HC. 2000. Real-time imaging of fluorescent flagellar filaments. J. Bacteriol. 182:2793–801. The video is reproduced with permission from Howard Berg’s website at the Rowland Insitute Harvard University (http://www.rowland.harvard.edu/labs/bacteria/movies/ecoli.php).
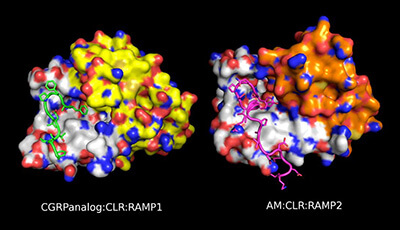
Receptor Activity-Modifying Proteins (RAMPs): New Insights and Roles: Supplemental Video 1
A supplemental video from the 2016 review by Debbie L. Hay and Augen A. Pioszak "Receptor Activity-Modifying Proteins (RAMPs): New Insights and Roles" from the Annual Review of Pharmacology and Toxicology.

Mangrove Sedimentation and Response to Relative Sea-Level Rise: Visual Abstract
A visual abstract for the 2016 review by C.D. Woodroffe K. Rogers K.L. McKee C.E. Lovelock I.A. Mendelssohn and N. Saintilan "Mangrove Sedimentation and Response to Relative Sea-Level Rise" from the Annual Review of Marine Science.

A Conversation with Maarten Schmidt - Other Work and Future Directions
Sandra Faber talks with Maarten Schmidt about his life and career including the discovery of quasars. Video 4 of 4.

A Conversation with Maarten Schmidt - The Discovery of Quasars
Sandra Faber talks with Maarten Schmidt about his life and career including the discovery of quasars. Video 1 of 4.

A Conversation with Maarten Schmidt - Early Life, Education, and Other Work
Sandra Faber talks with Maarten Schmidt about his life and career including the discovery of quasars. Video 3 of 4.

A Conversation with Maarten Schmidt - Using Quasars to Establish that the Universe Evolves
Sandra Faber talks with Maarten Schmidt about his life and career including the discovery of quasars. Video 2 of 4.

Group Affect
In this video Sigal Barsade of the Wharton School at the University of Pennsylvania and Andrew Knight of Olin Business School at Washington University discuss their article “Group Affect” which they co-wrote for the 2015 Annual Review of Organizational Psychology and Organizational Behavior. They explain how emotional contagion helps maintain group cohesiveness in a professional environment and how leaders can cultivate positive affect for better results.
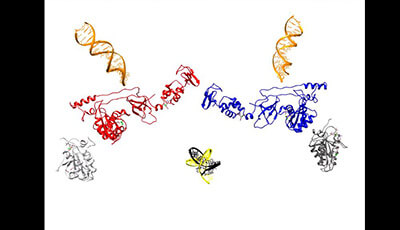
Retroviral Integrase: Then and Now: Video 1
A video from the 2015 review by Mark D. Andrake and Anna Marie Skalka "Retroviral Integrase: Then and Now" from the Annual Review of Virology.
Shown: Assembly of the prototype foamy virus (PFV) intasome (see Figure 3). This brief video portrays one possible scenario for how the PFV intasome may be assembled from PFV integrase (IN) monomers the predominant multimer found in solution. The complex is shown in ribbon representation with the inner subunits in red and blue. Viral DNA oligonucleotides are in orange ribbon ladder representation and the target DNA in yellow and black. The outer IN subunits only show the catalytic core domains and are colored gray. After the target DNA moves into place the view zooms in to one active site to show the location of active site metals (depicted as green spheres) and the DDE residues (shown in stick representation) that are required for metal binding. This video was generated using UCSF Chimera software [Pettersen et al. J. Comput. Chem. 25:1605–12 (2004)] with the coordinates of PDB 4E7K.
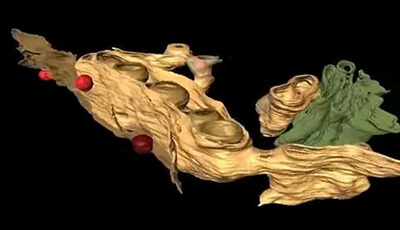
Flaviviridae Replication Organelles: Oh, What a Tangled Web We Weave: Video 1
A video from the 2015 review by David Paul and Ralf Bartenschlager "Flaviviridae Replication Organelles: Oh What a Tangled Web We Weave" from the Annual Review of Virology.
Shown: Morphology of dengue virus (DV) replication organelles. Electron tomography with 3D reconstruction reveals the continuity of endoplasmic reticulum membranes (yellow) and DV-induced vesicular invaginations. Budding of DV progeny virions (red) juxtaposed to vesicle openings is frequently observed. Golgi membranes are depicted in green. Adapted with permission from Welsch et al. Cell Host Microbe 5:365–75 (2009).
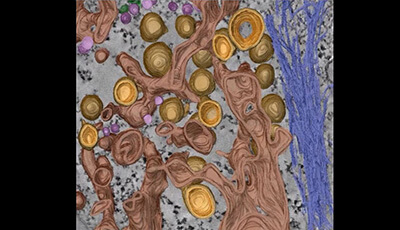
Flaviviridae Replication Organelles: Oh, What a Tangled Web We Weave: Video 2
A video from the 2015 review by David Paul and Ralf Bartenschlager "Flaviviridae Replication Organelles: Oh What a Tangled Web We Weave" from the Annual Review of Virology.
Shown: Morphology of hepatitis C virus (HCV) replication organelles consisting primarily of double-membrane vesicles (DMVs). Color-coded 3D surface model shows outer membranes of DMVs in light brown and inner membranes in orange. DMVs emerge as protrusions from the endoplasmic reticulum (dark brown) and are thus commonly found to be connected to this compartment. Small single-membrane vesicles are depicted in pink intermediate filaments in dark blue and Golgi membranes in green. Reproduced with permission from Romero-Brey et al. PLOS Pathog. 8:e1003056 (2012).
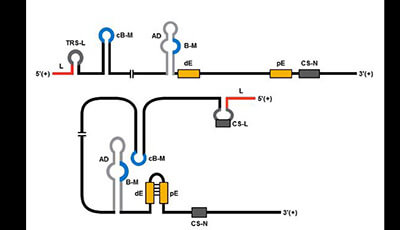
Continuous and Discontinuous RNA Synthesis in Coronaviruses: Video 1
A video from the 2015 review by Isabel Sola Fernando Almazán Sonia Zúñiga and Luis Enjuanes "Continuous and Discontinuous RNA Synthesis in Coronaviruses" from the Annual Review of Virology.
Shown: Model for the formation of genome high-order structures regulating N gene transcription. The upper linear scheme represents the coronavirus genome. The red line indicates the leader sequence in the 5′ end of the genome. The hairpin indicates the TRS-L. The gray line with arrowheads represents the nascent negative-sense RNA. The curved blue arrow indicates the template switch to the leader sequence during discontinuous transcription. The orange line represents the copy of the leader added to the nascent RNA after the template switch. The RNA-RNA interactions between the pE (nucleotides 26894 to 26903) and dE (nucleotides 26454 to 26463) and between the B-M in the active domain (nucleotides 26412 to 26421) and the cB-M in the 5′ end of the genome (nucleotides 477 to 486) are represented by solid lines. Dotted lines indicate the complementarity between positive-strand and negative-strand RNA sequences. Abbreviations: AD active domain secondary structure prediction; B-M B motif; cB-M complementary copy of the B-M; cCS-N complementary copy of the CS-N; CS-L conserved core sequence of the leader; CS-N conserved core sequence of the N gene; dE distal element; pE proximal element; TRS-L transcription-regulating sequence of the leader. Download a PowerPoint version of the animation.
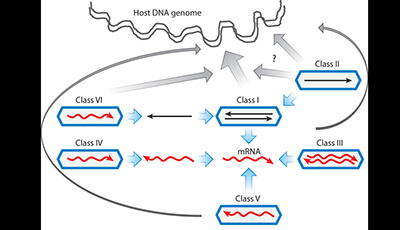
Deep Recombination: RNA and ssDNA Virus Genes in DNA Virus and Host Genomes: Video 1
A video from the 2015 review by Kenneth M. Stedman "Deep Recombination: RNA and ssDNA Virus Genes in DNA Virus and Host Genomes" from the Annual Review of Virology.
Shown: Schematic of different virus classes [after Baltimore 1971 (10)] and the integration of some viruses into cellular genomes. Black arrows represent DNA. Red arrows represent RNA. Blue hexagons represent virus capsids. Blue arrows represent virus replication mechanisms to produce mRNA for protein translation. Gray arrows represent integration into the host genome with the thickness of the arrow representing the gross frequency of recombination. The recombination arrow from Class VI viruses the retroviruses proceeds through a dsDNA intermediate. Recombination between ssDNA viruses (Class II) and dsDNA genomes may proceed through dsDNA (lower arrow) or directly as ssDNA (upper arrow) (55). Recombination between Class V and Class III viruses is probably mediated by cellular retrotransposons (see Figure 2). Download a PowerPoint version of the animation.
Deep Recombination: RNA and ssDNA Virus Genes in DNA Virus and Host Genomes: Video 2
A video from the 2015 review by Kenneth M. Stedman "Deep Recombination: RNA and ssDNA Virus Genes in DNA Virus and Host Genomes" from the Annual Review of Virology.
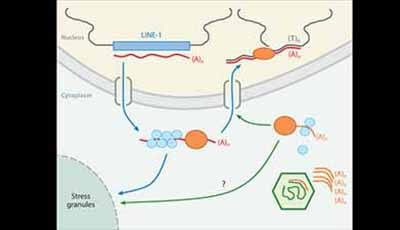
Shown: Proposed LINE-1-mediated mechanism of integration of RNA virus genes into host genomes. LINE-1 elements in the host genome (blue rectangle) are transcribed into mRNA (red line). The resulting mRNAs are exported to the cytoplasm where they combine with ORF1 (light blue circles) and ORF2 (orange ovals) the products of the two LINE-1 open reading frames. These complexes can be either relocated to stress granules or reimported back into the nucleus (blue arrows) where they can recombine with the host genome (dark gray lines) by reverse transcription. RNA viruses (green hexagon) infect the cell and produce mRNA (orange lines) that can bind to the ORF products of LINE-1 and be imported to the nucleus for reverse transcription or possibly relocated to stress granules (green arrows). Download a PowerPoint version of the animation.
Motors, Anchors, and Connectors: Orchestrators of Organelle Inheritance: Supplemental Video 1
A video from the 2015 review by Barbara Knoblach and Richard A. Rachubinski "Motors Anchors and Connectors: Orchestrators of Organelle Inheritance" from the Annual Review of Cell and Developmental Biology.
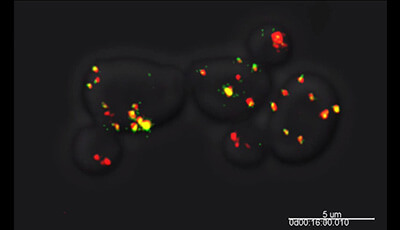
Shown: Inp1p: an organelle connector that tethers peroxisomes to the cell cortex. Budding yeast cells contain static (anchored) and mobile peroxisomes. Live-cell video microscopy was performed on wild-type cells expressing the peroxisomal matrix protein mCherry-PTS1 and Inp1p-3×GFP. Merged images of the red and green channels are presented. Static peroxisomes contain Inp1p and thus appear yellow whereas mobile peroxisomes are devoid of Inp1p and appear red.
The Hepatitis B Virus Receptor: Supplemental Video 1
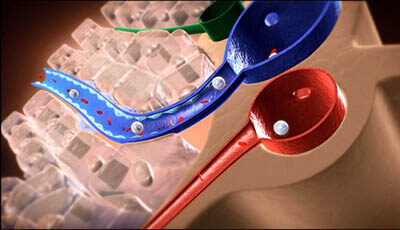
A video from the 2015 review by Wenhui Li "The Hepatitis B Virus Receptor" from the Annual Review of Cell and Developmental Biology.
